|
Molecular Mechanisms of Bacterial Pathogenicity |
The laboratory primarily focuses on three major areas of NIH-funded research, aiming to elucidate the molecular mechanisms of pathogenicity by Gram-negative and Gram-positive pathogens. We employ a multidisciplinary approach combining classical and modern techniques that include bacterial genetics, various biochemical methods, electron microscopy, electron cryo-tomography, X-ray crystallography, biophysics, mass spectrometry, cell-based assays, and rodent models of infection, hence providing a rich training environment to all trainees.
|
Virulence Determinants of Fusobacterium nucleatum
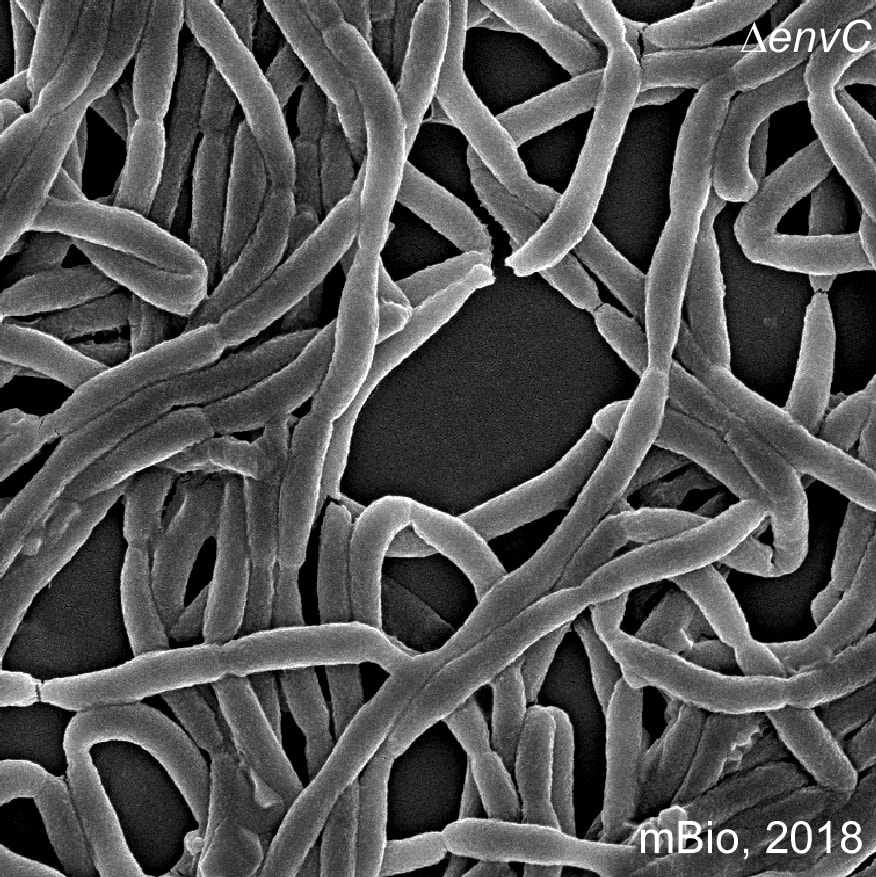
The Gram-negative pathobiont F. nucleatum is a key colonizer in the development of oral biofilms and well known for its association with human diseases including oral infections, preterm birth, and colorectal cancer. F. nucleatum has an inherent ability to interact with many early and late colonizers of the oral biofilms. It induces inflammatory responses and preterm birth in rodent models of infection, as well as promoting colorectal carcinogenesis and chemoresistance via induction of autophagy. Despite its pathogenic potential, we have limited knowledge about molecular mechanisms of fusobacterial virulence and associated factors. A major obstacle limiting progress is the lack of robust genetic tools and systematic investigations. We have tackled this problem with multiple complementary approaches including forward and reverse genetics, cryo-electron tomography, biochemical methods, and rodent models of infection. We are currently characterizing novel factors and pathways resulted from these experimental approaches that affect bacterial virulence and fitness, promotion of colorectal cancer, and induction of autophagy. Furthermore, we aim to understand how metabolism by F. nucleatum modulates its virulence potential. |
|
Molecular assembly on the cell surface of Gram-positive bacteria
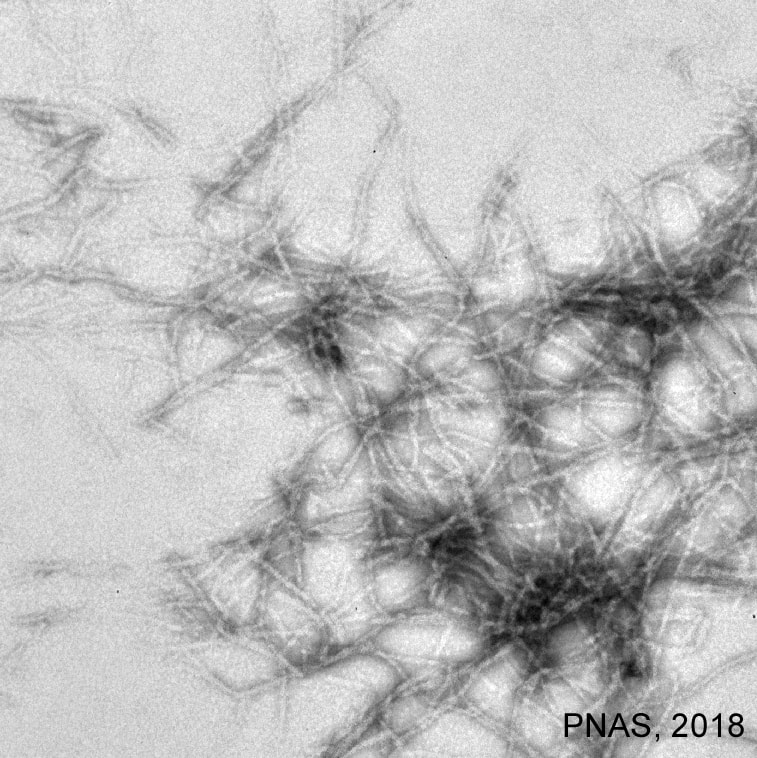
Gram-positive pathogens assemble on their surface covalently linked protein polymers known as pili or fimbriae that enable these bacteria to adhere to specific host tissues and initiate a pathogenic program. A typical pilus contains a major pilin forming the shaft and one or more minor pilin subunits. The heteromeric pilus is assembled by tandem transpeptidase enzymes called sortases. A pilus-specific sortase catalyzes the extension of pilus heteropolymers. The product of this cyclic polymerization is handed directly to the housekeeping sortase SrtA, which completes the assembly process by anchoring the resulting pilus polymer to the bacterial peptidoglycan. Several aspects of this biphasic mode of pilus assembly are not well understood. The current interests of the lab are to (1) examine the mechanism of pilus hijacking and polymicrobial interactions mediated by the major co-aggregation factor CafA in A. oris, (2) elucidate the mechanism of SrtA modulation of CafA spatial positioning and CafA-mediated coaggregation, and (3) examine the glycosylation mechanism of the cell wall anchored GspA that contributes to biofilm formation and membrane integrity. The commonality of these aspects is also being examined in Corynebacterium diphtheriae, Enterococcus faecalis, and Group B Streptococcus. Furthermore, we are collaborating with Drs. Robert Clubb (UCLA) and Jerzy Osipiuk (Argonne National Lab) to examine bacterial surface assembly at the atomic level. |
|
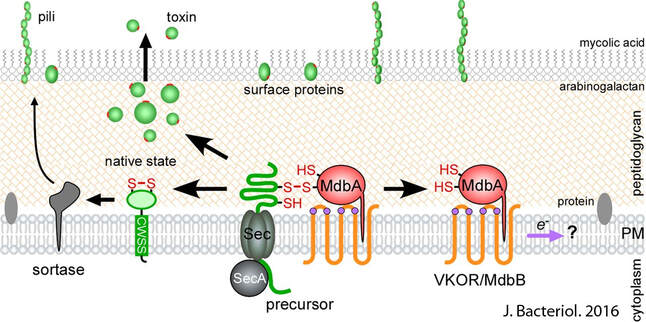
Post-translocational protein folding in Gram-positive bacteria
Many virulence determinants including pili and toxins are translocated across the cytoplasmic membrane by the Sec translocon in an unfolded state. How these proteins are maintained in a native state after translocation is not well understood in Gram-positive bacteria. We showed that the Gram-positive Actinobacterium A. oris employs a pair of membrane-bound oxidoreductase enzymes termed MdbA/MbdB as a general oxidative protein folding machine. Deletion of mdbA is lethal for cell growth, suggesting substrates of MdbA are involved in central cellular processes, including cell division and cell wall biosynthesis. The current focus of the lab is to determine the conservation of this pathway in other Gram-positive bacteria (C. diphtheriae and C. matruchotii), to reveal the molecular linkage between oxidative protein folding and cell wall biosynthesis, and to explore preventive strategies for dental caries and bacterial infections. |